Blastocyst grading is known to be highly subjective and without the use of time-lapse can result in an incorrect evaluation of collapsed or partially collapsed blastocysts. Artificial intelligence has the potential to make processes in IVF more precise, consistent and efficient in the future. This paper1 describes how deep learning (a subset of artificial intelligence) was used to design an algorithm to enable fully automatic grading of ICM and TE quality from time-lapse images. A similar approach has been used to estimate morphokinetic timings and PN count.
Algorithm for automatic grading of Inner Cell Mass and Trophectoderm
The group of Kragh et al. set out to determine if an automatic system based on deep learning can be helpful in the process of evaluating embryos, specifically when grading the inner cell mass and trophectoderm of blastocysts. This is a process which is currently manual and prone to subjectivity2. The deep learning algorithm described was developed based on a dataset of 780,000 images from time-lapse sequences of 8,664 embryos at the blastocyst stage -spanning from 90 hours post insemination to the time when the blastocyst had reached its maximum size. Of these sequences, 80% were used to train the neural network, and the remaining 20% were used to validate and test the algorithm.
The paper describes three main findings.
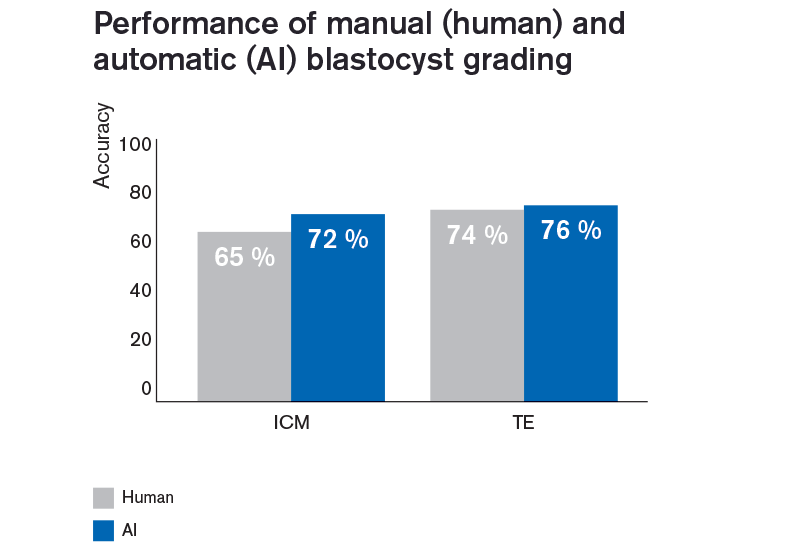
The algorithm performs better than humans when it comes to accurately grading blastocysts
Using the algorithm for inner cell mass and trophectoderm grading has the benefit of time savings, as the algorithm will provide the estimated grades instantly. However, in order for the algorithm to be relevant for daily use in the IVF laboratory, it is crucial that it also provides at least the same level of accuracy as manual grading by embryologists.
Testing the algorithm on human blastocysts cultured in the EmbryoScope time-lapse system shows a tendency for higher accuracy provided by the algorithm for ICM and TE grading.
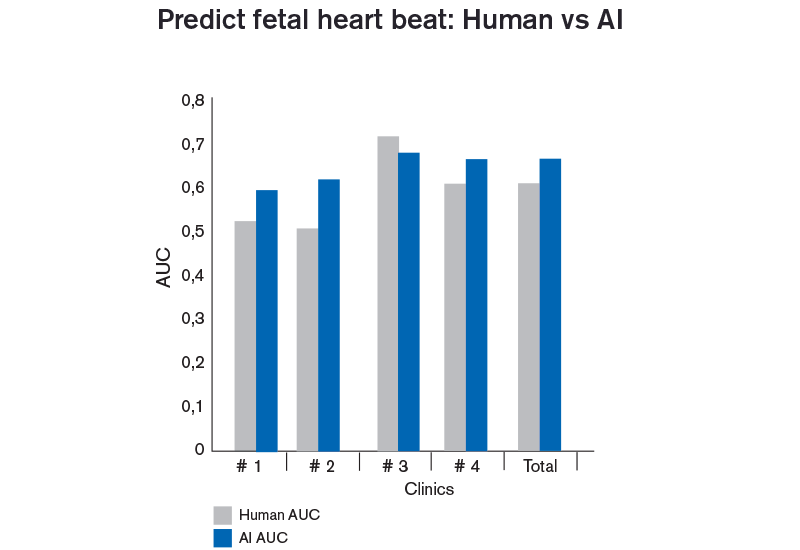
Predicting fetal heart beat
The study also tested the correlation of blastocyst grades with implantation (measured as fetal heart beat) by differentiating between top, good and poor quality blastocysts and testing against annotated clinical outcome from four different clinics. For this, the measure of performance was the area under the curve (AUC) of the receiver operating characteristic (ROC) and as shown below, the algorithm performed better than evaluation for most of the included clinics. Overall, the algorithm performed slightly better although without significance.
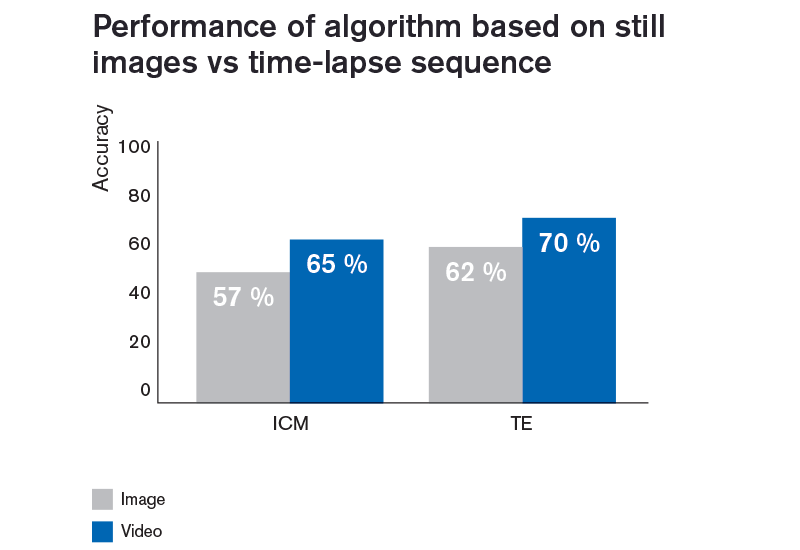
Inclusion of time-lapse information makes a positive difference
The inclusion of time-lapse sequences made a positive difference to the accuracy of the AI-based algorithm. Algorithms based solely on still images did not reach the same accuracy as an algorithm based on time-lapse sequences. This is shown in the graph below where “image” depicts algorithm designed based on still images whereas “video” depicts design based on time-lapse sequences. Combined with the above findings, this indicates that using time-lapse together with the AI-based algorithm may improve both laboratory workflow (by being able to retrieve the grading at a convenient time), efficiency (by providing instant and accurate gradings), consistency (as it will grade embryos in the same manner each time) and potentially also clinical outcome.
You can read the full study here.
Similar methods have been designed to estimate PN numbers and morphokinetic timings
PN number is commonly used as a check for correct fertilisation and many clinics benefit further from analysing morphokinetic parameters of embryos through the implementation of time-lapse systems.
With the use of AI-based methods that were trained on large datasets, both the PN number and the morphokinetic timings can be estimated very accurately. In a webinar, the author will show the performance of these algorithms when the system is set to provide estimates with a threshold of at least 90% accuracy.
Conclusion
The automatic algorithms for perform at least as good as the average embryologist for blastocyst grading and indirectly for predicting fetal heart beat as described above. Developing the algorithm based on time-lapse sequences lead to an improved accuracy compared to using only still images.
Training of deep learning algorithms is only based on raw image sequences and requires no prior knowledge of embryology. Thus, the algorithm learns by itself to extract the temporal and the morphological features that are most important for prediction of blastocyst grading.
It is important to note that in order to design and train a deep neural network, a substantial amount of data (in this case image sequences) is required. The data presented by Kragh et al. are based on image sequences from 8664 blastocysts which is the largest time-lapse dataset reported for development of AI-based embryo grading algorithms to date and similar methods were used to design algorithms for estimating PN numbers and morphokinetic timings.
The algorithms described here are all integrated into the Guided Annotation tool that works exclusively with the EmbryoScope time-lapse system. You can read more about Guided Annotation here.
More information in live webinar
Sign up to a live webinar with the author of the paper, Mikkel Fly Kragh, Tuesday, December 10th at 15 CET. Here, Dr. Kragh will present his and his co-authors’ work on integrating artificial intelligence into time-lapse and more specifically how they have automated blastocyst grading using AI.
References
1 Kragh, M.F., Rimestad, J., Berntsen, J., Karstoft, H. (2019): Automatic grading of human blastocysts from time-lapse imaging. Comput Biol Med, 115:103494
2 Richardson, A., Brearley, S., Ahitan, S., Chamberlain, S., Davey, T., Zujovic, L., … Raine-Fenning, N. (2015). A clinically useful simplified blastocyst grading system. Reprod BioMed Online, 31(4), 523–530.
Topics: Time-lapse
Written by Dr. Tine Qvistgaard Kajhøj
Tine did her PhD in the stem cell field. One of her responsibilities at Vitrolife is holding workshops where clinics both get started with and develop their skills in using time-lapse technology, in order to improve their results.
